# cellfinder-napari
[](https://github.com/napari/cellfinder-napari/raw/master/LICENSE)
[](https://pypi.org/project/cellfinder-napari)
[](https://python.org)
[](https://github.com/brainglobe/cellfinder-napari/actions)
[](https://codecov.io/gh/brainglobe/cellfinder-napari)
[](https://pepy.tech/project/cellfinder-napari)
[](https://pypi.org/project/cellfinder)
[](https://github.com/brainglobe/cellfinder-napari)
[](https://github.com/python/black)
[](https://pycqa.github.io/isort/)
[](https://github.com/pre-commit/pre-commit)
[](https://docs.brainglobe.info/cellfinder/contributing)
[](https://brainglobe.info/cellfinder)
[](https://twitter.com/brain_globe)
### Efficient cell detection in large images (e.g. whole mouse brain images)
`cellfinder-napari` is a front-end to [cellfinder-core](https://github.com/brainglobe/cellfinder-core) to allow ease of use within the [napari](https://napari.org/index.html) multidimensional image viewer. For more details on this approach, please see [Tyson, Rousseau & Niedworok et al. (2021)](https://doi.org/10.1371/journal.pcbi.1009074). This algorithm can also be used within the original
[cellfinder](https://github.com/brainglobe/cellfinder) software for
whole-brain microscopy analysis.
`cellfinder-napari`, `cellfinder` and `cellfinder-core` were developed by [Charly Rousseau](https://github.com/crousseau) and [Adam Tyson](https://github.com/adamltyson) in the [Margrie Lab](https://www.sainsburywellcome.org/web/groups/margrie-lab), based on previous work by [Christian Niedworok](https://github.com/cniedwor), generously supported by the [Sainsbury Wellcome Centre](https://www.sainsburywellcome.org/web/).
----
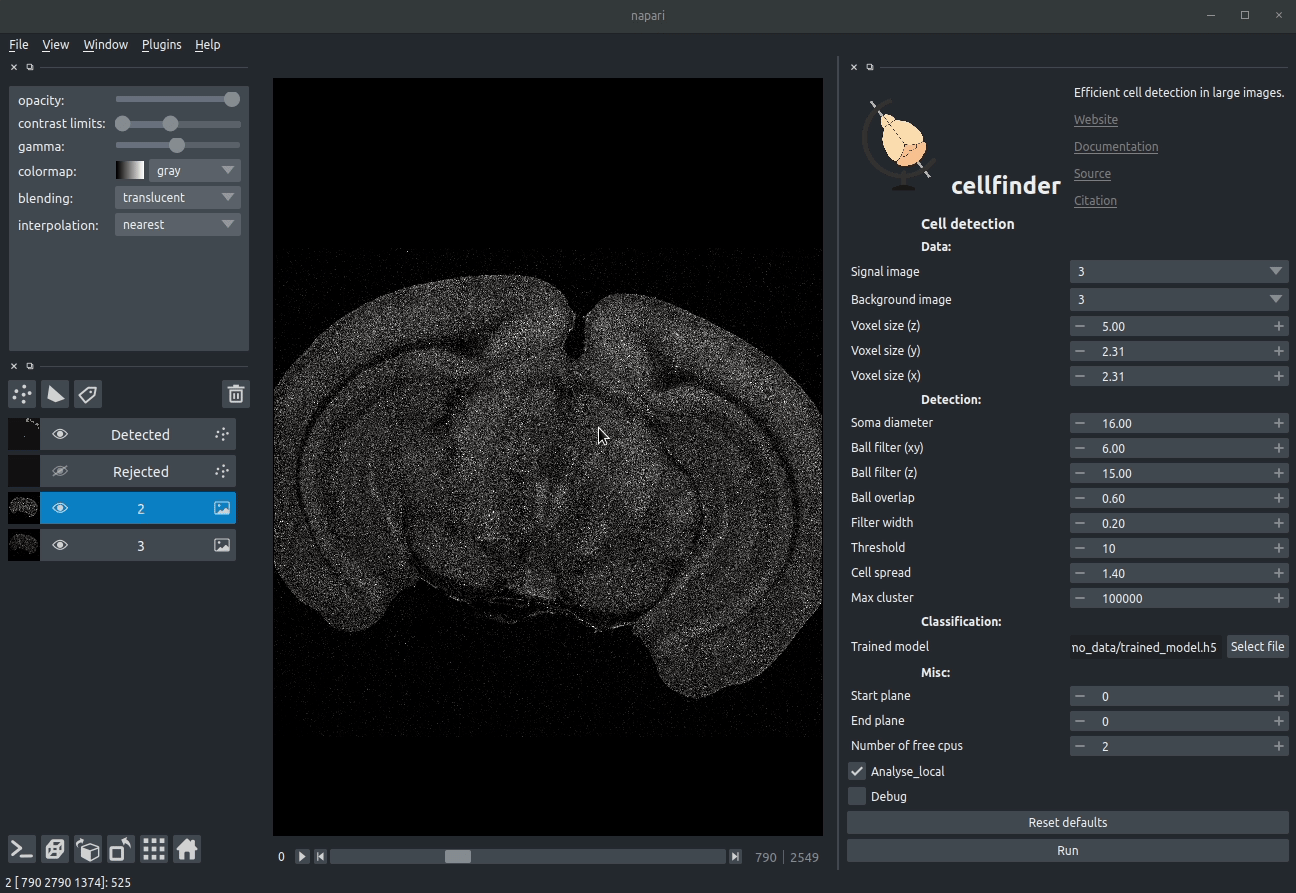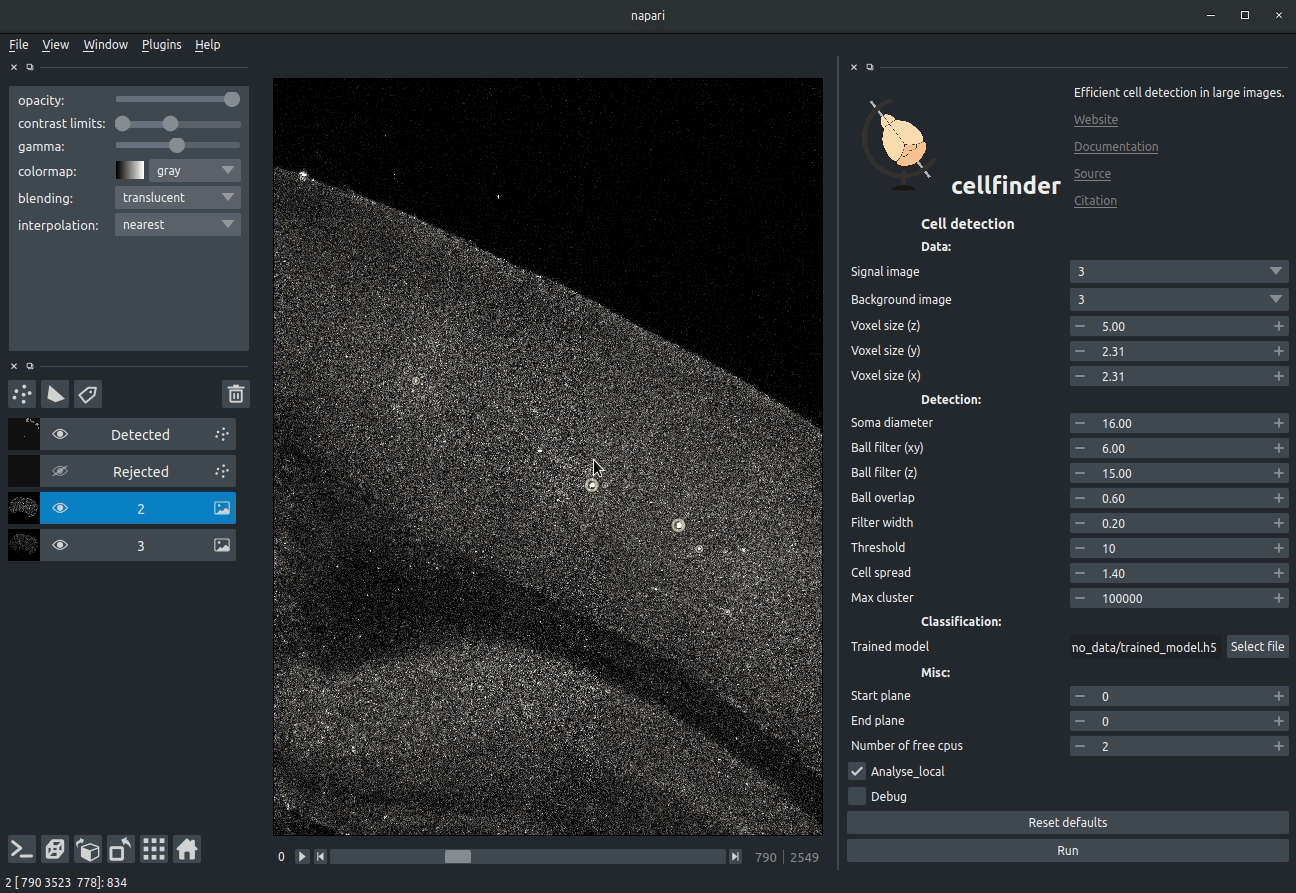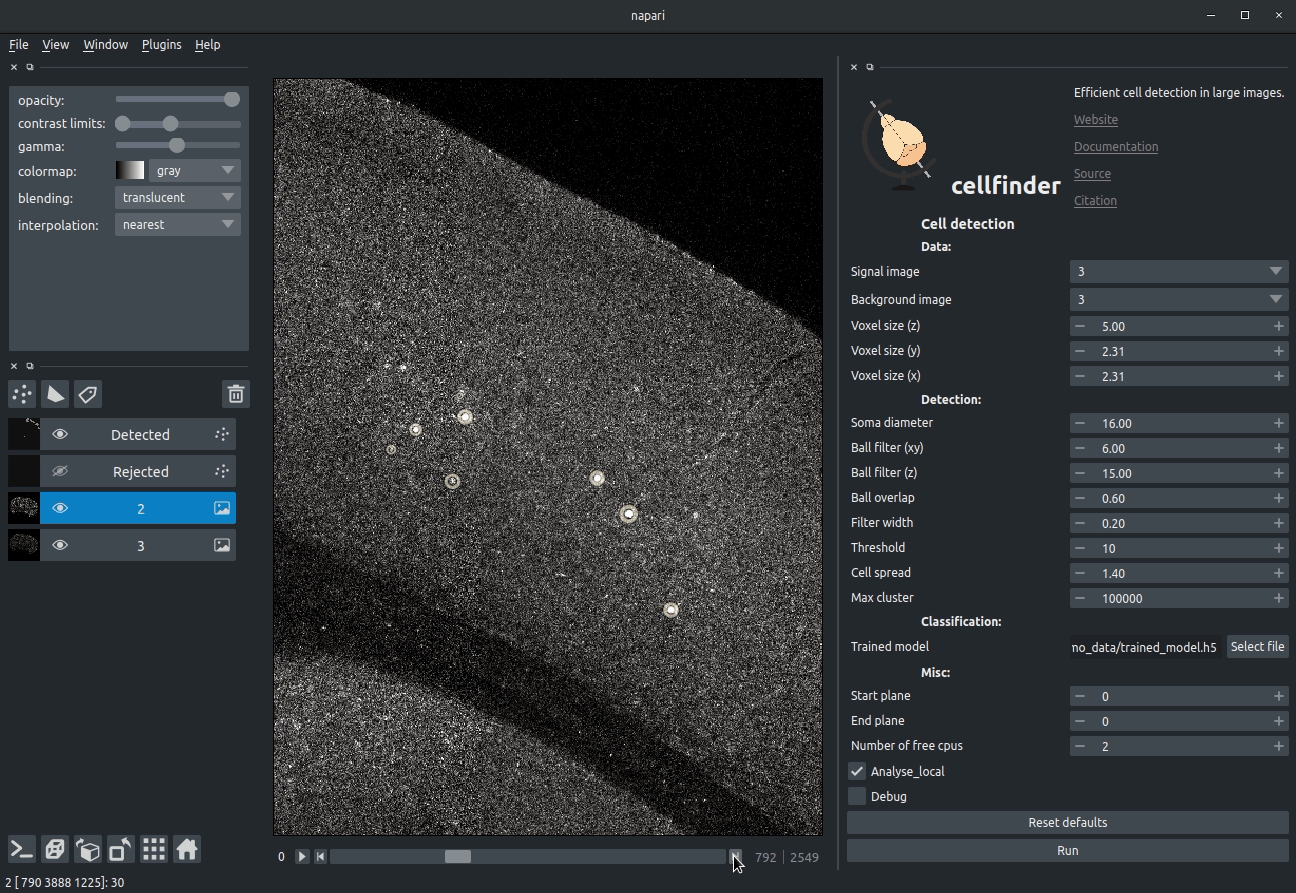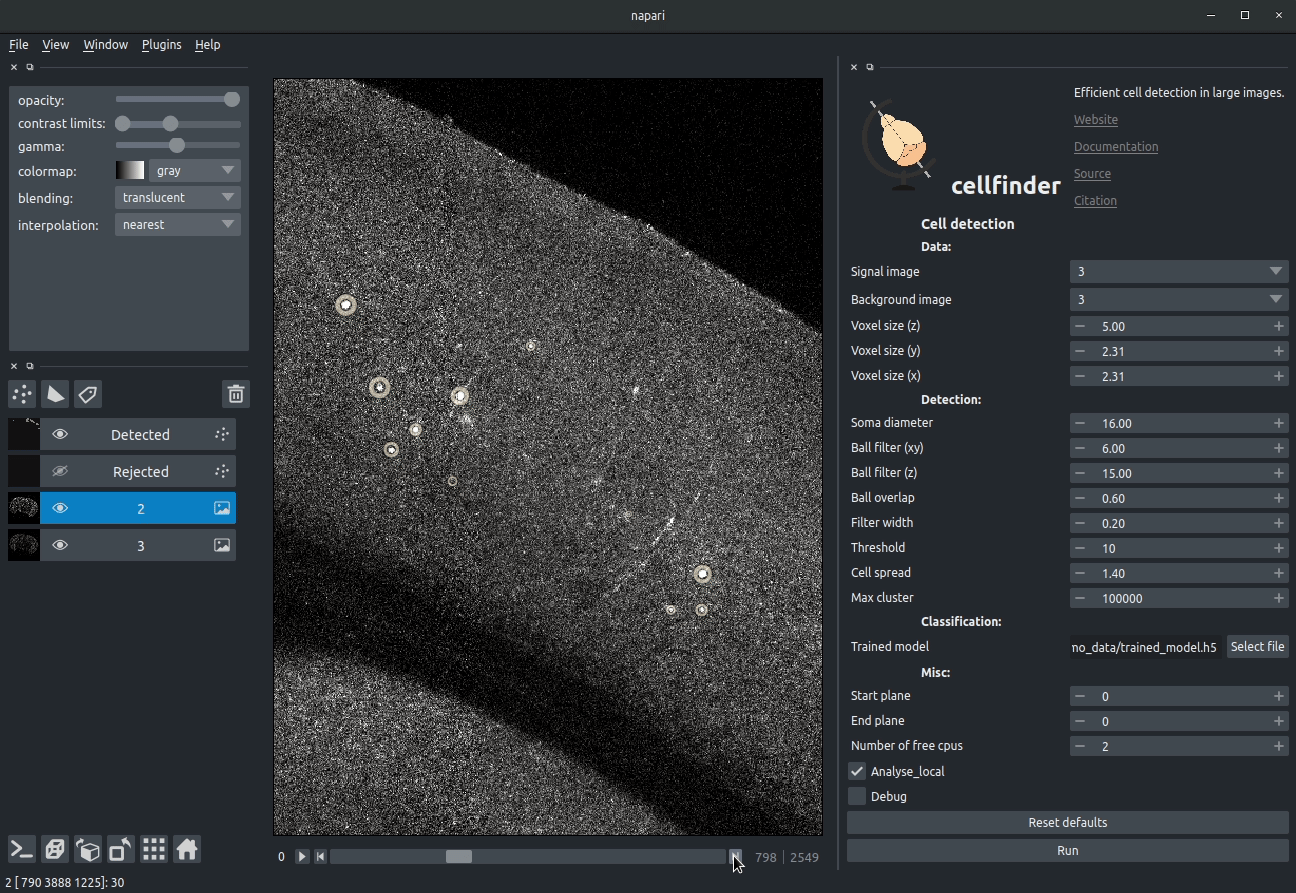
**Visualising detected cells in the cellfinder napari plugin**
----
## Instructions
### Installation
Once you have [installed napari](https://napari.org/index.html#installation).
You can install napari either through the napari plugin installation tool, or
directly from PyPI with:
```bash
pip install cellfinder-napari
```
### Usage
Full documentation can be
found [here](https://docs.brainglobe.info/cellfinder-napari).
This software is at a very early stage, and was written with our data in mind.
Over time we hope to support other data types/formats. If you have any
questions or issues, please get in touch [on the forum](https://forum.image.sc/tag/brainglobe) or by
[raising an issue](https://github.com/brainglobe/cellfinder-napari/issues).
---
## Illustration
### Introduction
cellfinder takes a stitched, but otherwise raw dataset with at least
two channels:
* Background channel (i.e. autofluorescence)
* Signal channel, the one with the cells to be detected:

**Raw coronal serial two-photon mouse brain image showing labelled cells**
### Cell candidate detection
Classical image analysis (e.g. filters, thresholding) is used to find
cell-like objects (with false positives):

**Candidate cells (including many artefacts)**
### Cell candidate classification
A deep-learning network (ResNet) is used to classify cell candidates as true
cells or artefacts:

**Cassified cell candidates. Yellow - cells, Blue - artefacts**
## Contributing
Contributions to cellfinder-napari are more than welcome. Please see the [contributing guide](https://github.com/brainglobe/.github/blob/main/CONTRIBUTING.md).
## Citing cellfinder
If you find this plugin useful, and use it in your research, please cite the preprint outlining the cell detection algorithm:
> Tyson, A. L., Rousseau, C. V., Niedworok, C. J., Keshavarzi, S., Tsitoura, C., Cossell, L., Strom, M. and Margrie, T. W. (2021) “A deep learning algorithm for 3D cell detection in whole mouse brain image datasets’ PLOS Computational Biology, 17(5), e1009074
[https://doi.org/10.1371/journal.pcbi.1009074](https://doi.org/10.1371/journal.pcbi.1009074)
**If you use this, or any other tools in the brainglobe suite, please
[let us know](mailto:code@adamltyson.com?subject=cellfinder-napari), and
we'd be happy to promote your paper/talk etc.**
---
The BrainGlobe project is generously supported by the Sainsbury Wellcome Centre and the Institute of Neuroscience, Technical University of Munich, with funding from Wellcome, the Gatsby Charitable Foundation and the Munich Cluster for Systems Neurology - Synergy.
<img src='https://brainglobe.info/images/logos_combined.png' width="550">




